 Epigenetic alterations of histones can be passed on to daughter cells. [1]
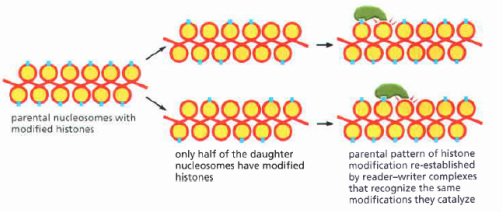
Epigenetic alterations of histones can be passed on to daughter cells. [1]
One type of inheritance, epigenetic inheritance, is a way for the cells to pass on the histone code protein structure or the covalent modifications associated with the histone proteins. This type of inheritance is unlike typical genetic inheritance in which a permanent change in DNA is passed on through cell generations. The packaging of DNA allows for the complexity associated with multicellular organisms; cell memory keeps the chromatin states persistent from generation to generation [1]. Nucleosomes that are part of the parental chromosomes have modifications on them. When the parent cell’s DNA is replicated, only half of the modifications are going to be present on each new set of DNA (a set of modifications on each strand) because of semi-conservative DNA replication. When the parent cell is divided into two daughter cells, only half of their nucleosomes are going to have the histone modifications [1]. Fortunately, the cell “remembers” the modifications that it has had in the past due to positive feedback loops that were created by the stimuli that caused the modifications in the first place [1]. This memory leads to the cell using the fore mentioned code-reader/code-writer complexes to reinstate the code onto these daughter nucleosomes.
 The location of chromatin in the nucleus influences gene expression.[1]
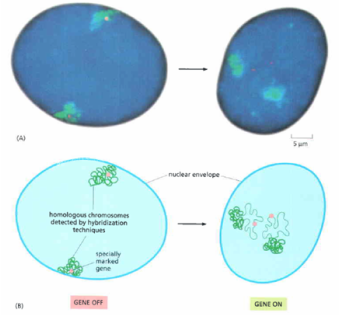
The location of chromatin in the nucleus influences gene expression.[1]
The nucleus has a heterogeneous environment. The chromosomes here are located in discrete environments; however, the movement of chromatin to various locations can alter the expression status of genes. The areas to which the chromatin moves are known as “nuclear neighborhoods”. These areas can be associated with gene expression or repression and therefore have a concentration of gene expression-related proteins or repressor proteins [1].
 There are various types of regulation that can occur through gene regulatory proteins. [1]
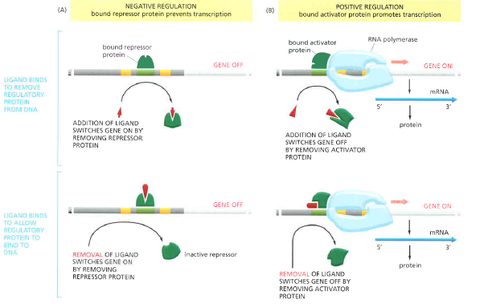
There are various types of regulation that can occur through gene regulatory proteins. [1]
Another way to regulate the activity in genes is through gene regulatory proteins, which are part of genetic switches along with a particular DNA sequences. These proteins are able to locate specific sequences on long DNA helices within the chromosome. Once they find a sequence, they bind to that portion of the DNA in order to turn genes on or off depending on the signal received [1]. In order to bind to specific DNA sequences, the gene regulatory proteins must be able to distinguish between the various patterns created on the DNA double helix by hydrogen bond donors, hydrogen bond acceptors, and hydrophobic patches in the major and minor grooves [1]. Many prokaryotes have genetic switches that can rather simply be turned on or off through positive or negative regulation. It is also helpful that in prokaryotes genes that need to be expressed together to work toward a common goal are usually located next to one another on the DNA sequence, in what is known as an operon [1]. Positive regulation can occur in two ways. First, the addition of a ligand can turn the gene off by causing an already bound activator protein to fall off of the DNA. A second scenario is that the activator protein needed to have the ligand in order to bind to the DNA; removal of the ligand would now be the cause of the activator protein to come off of the DNA. Both of these processes turn the genes off. Negative regulation, on the other hand, works to turn genes on. The addition of a ligand here means that the repressor protein will be removed from the DNA or the removal of a ligand from an already bound repressor will cause its release. Either way the gene is turned on (Alberts et. al., 2008, p. 436). Because of increased complexity and higher order packaging of DNA in higher organisms, simple on and off switches are not going to be able to work [1]. However, eukaryotes have evolved the ability of the use of more complicated switches that involve many more gene regulatory proteins and much more DNA. This complexity allows eukaryotic cells the advantage of responding to many signals versus the very few in prokaryotic cells [1].
 There are six main ways that repressor proteins work in order to halt gene expression. [1]
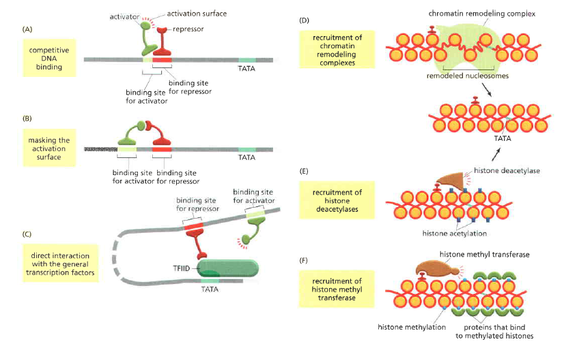
There are six main ways that repressor proteins work in order to halt gene expression. [1]
Gene activator proteins are a useful way to promote the expression of specific genes in eukaryotes. These proteins bind to specific DNA sequences on the chromosomes to promote transcription of certain genes. Each activator consists of a recognition domain (to find a specific DNA sequence) and an activation domain (to increase transcription) in the least. These activators work to attract gene regulatory proteins (general transcription factors) and other transcription machinery, such as the Mediator and RNA polymerase, to promoter sequences in DNA to allow the initiation of transcription. They may also directly modify the regulatory proteins or other transcription machinery, which will in turn cause the modification of the chromatin [1]. None of these three can adequately stay atop the DNA while it is packaged up in nucleosomes. The activator proteins can alter this packaging by causing chromatin modifications. They attract reader-writer complexes, chromatin remodeling complexes, and chaperones. By creating a less tightly compacted form of the chromatin, the transcription machinery is allowed better access to the DNA thus allowing better assembly at the promoter DNA regions. The changes that these activators cause must be reversed at some point in time. Some of the modifications last longer than others because some genes must be immediately switched off after their transcription [1]. The activators typically cause acetylation of the histones to loosen the chromatin, but they also cause the removal of nucleosome structures completely. When the nucleosomes are removed, they are re-added to the DNA behind the replication machinery. After transcription of the desired sequence occurs, the cell can recruit HDACs and methyl transferases to deacetylate and methylate the histones, respectively, in order to make the genes resistant to continual transcription [1]. Along with gene activator proteins, there are also gene repressor proteins, which act to inhibit transcription of a gene. There are six ways that repression proteins work. The first way is to directly compete with an activator protein to bind to the DNA. This occurs when the regulatory sequences are partially overlapping. A second way that these proteins can operate is through the binding of the repressor protein to the activation portion of the activator. In this situation, both proteins have recognized and bound to their respective regulatory DNA sequences. Thirdly, the repressor proteins may act by blocking the binding of transcription factors to either DNA or one another. They can also attract reader-writer complexes in order to further condense the chromatin to prevent the access of transcription machinery. Repressors may also attract HDACs or histone methyl transferases to the sites to cause chromatin compaction. Repression of genes is an important mechanism especially in eukaryotes because of the complex developmental processes that must occur at precise times [1].
Without the structure of chromatin, DNA could not fit inside of the nucleus.
Chromatin, Its Structure and Regulation
Edited by Cheryl Ann Fleming and Ashley Lauren McPhee
